Journal of Environmental and Agricultural Sciences (JEAS). Akbarzai et al., 2022. Volume 24(1&2): 18-25
Open Access – Research Article
AMMI Analysis of Yield Stability and Adaptability in Barley: A Case Study of Afghanistan
Darya Khan Akbarzai 1,*, Gull Rahman Akbari 2, Lina Mohammadi 1, Mohammad Navin Safi 1,
Abdulhaq Farhang 1
1International Center for Agricultural Research in the Dry Areas (ICARDA), Kabul, Afghanistan
2Agriculture Research Institute of Afghanistan (ARIA), Kabul, Afghanistan
Abstract: Barley, an important cereal crop in Afghanistan, has low productivity. To identify high yielding and stable genotypes, an evaluation of nine barley inbred lines and a national check was conducted for yield and its stability at 6 diverse environments in Afghanistan. Experiments were designed to determine genotype (G) × environment (E) interaction (GEI) effect on grain yield using AMMI model and to identify the high yielding and stable barley genotypes. The main effect of G and GEI were highly significant (P<0.01) on grain yield. The E, G, and GEI accounted for 75.0%, 7.5% and 8.2% of variation in grain yield. Based on the AMMI stability parameters, line G9 was the most stable lines across environments with above average grain yield and will be recommended as a candidate for the consideration of the Varietal Release Committee in Afghanistan.
Keywords: Barley, GE interaction, AMMI model, Grain yield, Stability
*Corresponding author: Darya Khan Akbarzai, email: d.akbarzai@gmail.com
Cite this article as
Akbarzai, D.K., G.R. Akbari, L. Mohammadi, M.N. Saif and A. Farhang. 2022. AMMI analysis of yield stability and adaptability in barley: A case study of Afghanistan. Journal of Environmental & Agricultural Sciences. 24(1&2): 18-25. [Abstract] [View Full-Text] [Citations] 
1. Introduction
Barley (Hordeum vulgare) is a major cereal crop cultivated under diversified agroclimatic conditions, from temperate to arid and semi-arid regions of the world, including Afghanistan (Kaur et al., 2022). Globally it is ranked fourth in the cereal production, after maize, rice and wheat (Yirgu et al., 2022). Barley belongs to cereal family Poaceae and three types of cultivated barley are Hordeum vulgare (a six-rowed type), Hordeum distichum (a two-rowed type) and Hordeum irregulare (the least cultivated type) (Bedada et al., 2014; Russell et al., 2016). Barley was domesticated 10,000 years ago from wild relative Hordeum spontaneum in Israel-Jordan region (Badada et al., 2014; Badr et al. 2000; Zohary, 2017). As it is a cool-season crop, it can also be successfully grown from an altitude of sea level to more than 3000 MSL and well adopted in stressed environment where soil erosion, drought and frost are the main problems for several crops (Fana et al., 2018). Barley is a multipurpose crop with several economic value and utilization (Newman et al., 2019; Sharma et al., 2022; Zhou, 2010) and have nutritive components, comparable with maize and other cereals (Lyu et al., 2022; Meints et al., 2021; Obadi et al., 2021). Barely has been used primary for animal feed (Perera et al., 2022; Raud et al., 2021; Sakellariou and Mylona, 2020), hay and ethanol production (Diaz et al., 2022; Soufan and Al-Suhaibani, 2021; Tse et al., 2021).
Russian Federation, Germany, France, Ukraine, Australia, Canada, Spain, Turkey, United Kingdom and USA are top barely-growing countries of the world (Ullrich, 2014; Mittal, 2022) with total production of 157.0 million tons from 51.1 million hectares with an average yield of 3.31 t/ha, while in Afghanistan, a total of 127.7 thousand Mt barley produced in the area of 86.0 thousand hectares with productive of 1.48 t/ha (FAO, 2020; Tricase et al., 2018). The barely production showed negative trend (from 514 down to 301.8 thousand Mt) including area of production and productivity (by 58.7 thousand ha and 0.47 t/ha) between 2013 and 2016, respectively. The low production and productivity are strictly associated with the unavailability of improved barley variety with wide adaptability and stability across the country and negative effect of several biotic and abiotic factors as drought, frost, diseases and poor soil (Alasti, et al., 2020; Chapagain and Good, 2015; Cossani et al., 2010; Schils et al., 2018). USDA (2018) reported that import (barley grain) has been increased since 2014 to meet the domestic demand due to decreasing in production.
Crop production, in drier regions of the world, including Afghanistan, is highly dependent on utilization of improved variety and adaptation of improved crop production practices, efficient soil and water management and agrochemical practices (Akbarzai et al., 2021a; Belachew et al., 2022; UnNisa et al., 2022; Zhang et al., 2016). Hence, there is need for the development of improved and high yielding cultivars, with absolute stability and wide adaptation, to fill the current production gape and farmers requirement in Afghanistan (Akbarzai et al., 2021b). Therefore, multi-environment trials (MET) were conducted in plant breeding program to identify high yielding and widely or specifically adapted genotypes for different environments. MET ensure the evaluation of number of genotypes across the years and locations and has two main goals (1) to identify the favorable environments and (ii) to identify the high yielding genotypes (Vaezi et al., 2017). The selection of superior genotypes in METs generally results in genotype-by-environment interactions that frequently make difficult the interpretation of results obtained and reduce efficiency of selection (Annicchiarico and Perenzin, 1994).
The performance of genotypes arises from the interaction of genotype and environment (GEI). The environmental factors may be characterized by biotic and abiotic stresses. Statistical techniques have been developed for the analysis and interpretation of GEI from MET where the commonly used statistical methods for analyzing GEI are the additive main effects and multiplicative interaction (AMMI) model (Ebdon and Gauch, 2002). The AMMI is an effective model for analysis of multi-environment yield trials (MEYTs) where it describes a large portion of the GE sum of squares uniquely partitioning into interpretable principal components (IPC) (Adil et al., 2022; Ebrahimnejad and Sabouri. 2018; Kebede and Getahun, 2017; Zhang et al., 2020). The main objectives of this study were (i) to apply AMMI to analyze the GE interaction effect on grain yield (ii) to identify the high yielding and stable barley genotypes within test environments.
2. Materials and Methods
2.1. Experimental Conditions and Genetic Material
This study is based on the experiments conducted in six environments—three locations (Mazar, Baghlan and Bamyan) over three years (2016-17, 2017-18, 2018-19). The genetic materials (Table 1) used in this experiment to determine the yield performance of barley inbred lines including one national check (improved variety) at Dahdadi research farm in Mazar (36° 39 25.4 N, 66° 57 39.9 E, 398 m asl, average annual precipitation 282 mm), Posi-Shan research farm in Baghlan (36° 09 N, 68° 64 E, 564.9 m asl, average annual precipitation is 268.4 mm) and at Molaghalam research farm in Bamyan (at 34 °43 N, 67° 49 E, 2550 m asl , the average annual precipitation is 321 mm) during 2016, 2017 and 2018 cropping season.
The experimental design was a randomized complete block design with three replications at each site across three years. A plot consisted of four 5-m rows with 0.20 m row-to-row distance and 6 rows per plot, using the four central rows for evaluation. Weed management and pest and disease control were carried out according to the recommended cultural practices for Barely production by the Agricultural Research Institute of Afghanistan (ARIA). Sowing was done during November – December, and the crop was harvested during June – July. The grain yield was obtained from the central four rows (3.2m²) plot area for all the trials to remove the border effects. The yield was converted to ton per hectare for statistical analysis.
2.2. Statistical analyses
AMMI model was used to assess the genotype by environment interaction (GEI), adaptability and stability of genotypes in the tested environments. The AMMI model integrates the standard ANOVA with principal components (PC) analysis as described by Zobel et al., 1988. The AMMI model was applied to determine the effect of genotype (G), environment (E) and genotype by environment interaction (GEI).
During analyses, the combined among one to three locations and the three years were considered as an environment giving rise to six environments: Mazar, 2016 (E1), Mazar, 2017 (E2), Baghlan, 2017 (E3), Mazar, 2018 (E4), Baghlan, 2018 (E5), Bamyan, 2018 (E6). All obtained data were subjected to analysis using GenStat software (VSN Inc. 2015).
Table 1. The names and pedigree of nine tested barely lines including check and the environments

3. Results and Discussion
3.1 AMMI Model Analysis
The result of AMMI analysis for grain yield of 10 barely inbred lines and 6 environments are given in Table 2. The main effect of genotypes and genotype × environment interaction (GEI) was highly significant (p<0.01). The result further indicated that environments (E), genotypes (G) and genotype × environment interaction (GEI) accounted for 75.01%, 7.5% and 8.16% of the total variation, respectively. Furthermore, results showed that the environment is the predominant source of variation and grouping the environments may provide a clearer understanding of the GEI. There were significant variations in environmental conditions. Moreover, the performance of genotypes significantly differed across the tested environments as indicated by a higher GEI component than the genotype component (Shukla et al. 2015). Other researchers also reported a higher percentage of G×E relative to the genotype and recommended the stability analysis and partitioning of GEI into its component (Amiri et al., 2013; Homma 2015; Erdemci 2018).
Table 2. AMMI analysis of variance for grain yield (t/ha) of the 10 barely genotypes tested over six environments (combination of three locations and three years).
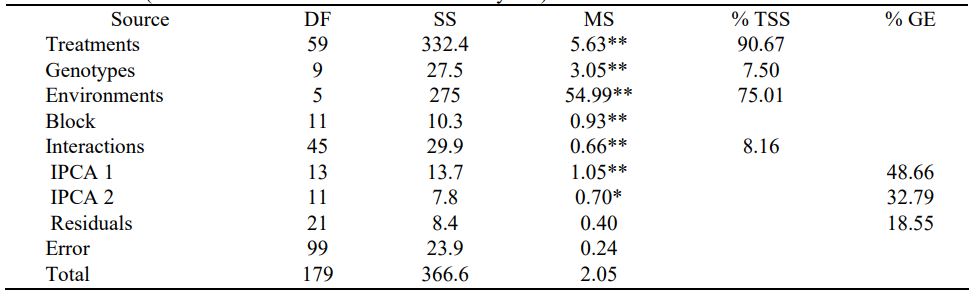
*, ** Significance at 5 and 1 percent probability levels, respectively.
Table 3. Mean Grain yield (t/ha), first and second Interaction Principal Components Analysis (IPCA), AMMI stability Value (ASV) of the 10 barley genotypes over the six environments (combination of three locations and three years)
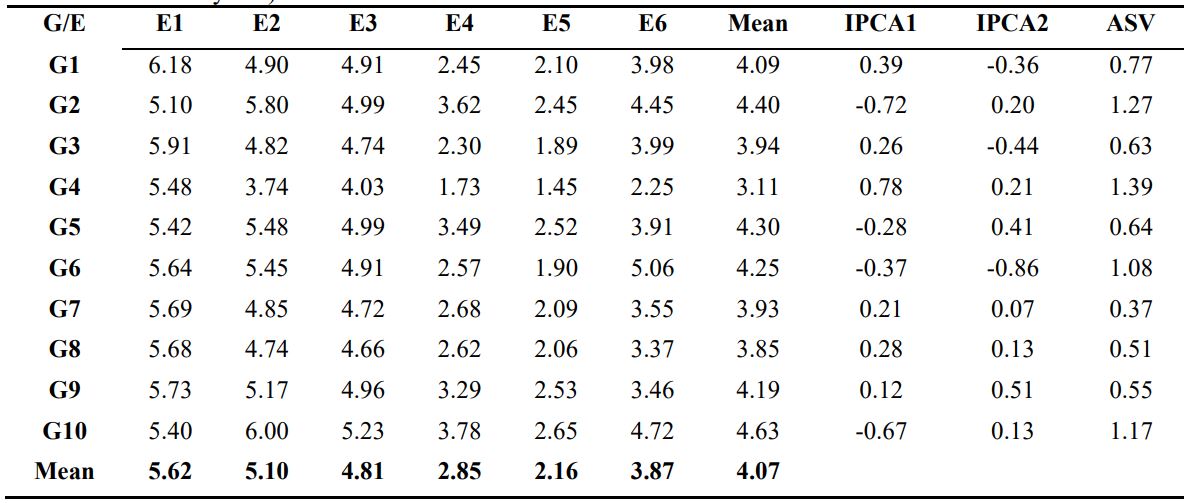
G: genotype; E: environment
The AMMI estimates the main effect of genotypes and multiplicative interaction as principal components, IPCA1 and IPCA2, found suitable to model to predict the GEI in grain yield (Gauch and Zobel, 1996). In our study, the IPCA1 and IPCA2 were significant and explained 48.66% and 32.79% of the total variation in G×E interaction. Therefore, the two PCAs axes presented 81.45% of the interaction sum of squares (GEI) with 18.55% residual contribution. The result was consistent with the findings of other researchers (Mortazavianc et al., 2014; Fana et al., 2018) from the study on barley.
The mean value of the genotype and the environment are given in Table 3. The environment means for grain yield (mean of genotypes) varied from 2.16 t/ha at E5 to 5.62 t /ha at E1. The E1, E2 and E3 environments had higher than average (4.07 t/ha). The mean of grain yield of tested genotypes varied from 3.11 t/ha for G4 to 4.63 t/ha for G10. G10 yielded higher than average in 4 out of 6 environments and 60 % of genotypes yielded higher than the overall genotypic mean with the contribution of 6 barley lines viz. G10, G2, G5, G6, G9 and G1 constantly yielded higher than average (4.07 t/ha). Ahmadi et al. (2012) reported four high-yielding genotypes (G1, G14, G11 and G2) out of 18 barley genotypes based on mean performance across tested environments; Vaezi et al. (2017) grouped the barley genotypes as high (G9, G2 and G5) and low (G6, G8 and G10) yielding in compression to genotypes mean (2.09 t/ha) where the genotypes mean was in the range of 1.86 t/ha – 2.30 t/ha.
3.2. AMMI stability value (ASV)
Purchase et al. (2000) introduced, ASV, as a measure of stability of the genotype based on two principal components under AMMI Analysis. ASV is the distance of the varieties from point zero of the scatter diagram (IPCA1 vs. IPCA2). Therefore, lower score of ASV and IPCA1 presenting to high stability of genotypes. The genotypes G7, G8 and G9 are identified as three most stable genotypes due to their low value of ASV (0.37 – 0.55), where the G4, G2, G10 and G6 are recognized as unstable genotypes (ASV: 1.08 – 1.39) (Table 3). Amiri et al. (2013) also grouped the wheat genotypes based on ASV analysis as stable and unstable; Vaezi et al. (2017) reported and identified the 5 barely line as stable one with a small ASV score and less interaction with environments while the other 5 lines had high ASV score and detected as unstable lines.
3.3. AMMI biplot analysis
Based on the AMMI biplot which estimates the genotype stability and adaptability differences in related environments by using IPCA vs IPCA2. Therefore, the AMMI biplot is presenting the yield difference of 10 genotypes in 6 environments in Fig. 1. A stable genotype has a value closer to the origin of the axis (IPCA1) with a small contribution to the interaction (Gauch, 1992). The variation due to environment is higher than the genotype in both main effects and interactions (IPCA1).

Fig. 1. The AMMI biplot (IPCA1 vs mean) for barley yield (t/ha) of 10 genotypes across six environments (combination of three locations and three years).
Among the genotypes, G9 is the most stable genotype with above average yield and wide adaptability while G4 and G2 were the unstable genotypes although G2 had mean over the average mean. In the biplot, the genotype G1, G2, G5, G6, G9 and G10 had higher average yields and adapted to favorable environments, while the genotypes G3, G4, G7 and G8 were adapted to poor environment. According to environmental index value (negative and positive), the environments were separated as rich (E3), medium (E1 and E2) and poor (E4, E5 and E5). These results are in agreement with the findings of earlier reports for rice (Islam et al., 2014); wheat (Erdemci et al., 2018) and barley (Fana et al., 2018).
4. Conclusion
The result indicated that the AMMI analysis is the best statistical method for the identification of stable and wide and specific adapted genotypes in MET. The AMMI analyses indicated, the environment has a high effect on the grain yield of barley rather than the effect of GEI and genotype. Therefore, the stability analysis is necessary to determine the stable and wide adapted genotype in tested environments. According to AMMI biplot analysis, the G5 and G9 were the most stable and wide (G9) and specific (G5) adapted genotypes with grain yield mean of above average and could be considered for further release as variety.
Competing Interest Statement: Authors declare that there is no conflict of interests arising from this study.
List of Abbreviations: AMMI: the additive main effects and multiplicative interaction, ANOVA: Analysis of variance, Asl: Above sea level, ASV: AMMI Stability Value, DF: degree of freedom, E: environment, FAO: Food and Agriculture Organization of the United Nations, G: Genotype, GEI: Genotype by environment interaction, ICARDA: International Center for Agricultural Research in the Dry Areas, IFAD: International Fund for Agricultural Development, MET: Multi-environment trials, PCA: Principal Components Analysis, USDA: United States Department of Agriculture.
Competing Interest Statement: Authors declare that there is no conflict of interests arising from this study.
Author’s Contribution: Conceptualization, D.K.A. L.M. and G.R.A.; Data curation, D.K.A. and L.M.; Formal analysis, D.K.A.; Funding acquisition, IFAD; Investigation and Methodology, D.K.A., G.R.A., L.M., M.N.S. and A.F.; Resources, ICARDA and ARIA; Validation, M.S. and NS; Visualization, D.K.A.; Writing – original draft, D.K.A.; Writing – review & editing, M.S., R.M.R., H.M., and A.H.
Acknowledgment: The authors would like to thank Prof. Murari Singh, from Concordia University for his kind suggestions and review. The authors also acknowledge the Agriculture Research Institute of Afghanistan (ARIA) for providing the necessary support and facilities.
References
Adil, N., S. H. Wani, S. Rafiqee, S. Mehrajuddin, N. R. Sofi, A. B. Shikari, A. Hussain, F. Mohiddin, I. A. Jehangir and G. H. Khan. 2022. Deciphering genotype× environment interaction by AMMI and GGE biplot analysis among elite wheat (Triticum aestivum L.) genotypes of Himalayan region. Ekin J. Crop Breed. Genet. 8: 41-52.
Ahmadi, J., B. Vaezi and M.H. Fotokian. 2012. Graphical analysis of multi-environment trials for barley yield using AMMI and GGE-biplot under rain-fed conditions. J. Plant Physiol. Breed. 2(1): 43-54.
Akbarzai, D.K., O.J. Mangal, S. Nisar and L. Mohammadi. 2021 a. On-farm assessment of productivity of improved varieties of wheat. J. Innov. Agri. 8(3): 11-16.
Akbarzai, D.K., S. Nisar and L. Mohammadi. 2021 b. Genotype × Environment interaction studies in lentil under Afghanistan environments. J. Innov. Agric. 8(2): 39-46.
Alasti, O., E. Zeinali, A. Soltani and B. Torabi. 2020. Estimation of yield gap and the potential of rainfed barley production increase in Iran. J. Crop Prod. 13: 41-60.
Amiri, E., Z. Farshadfar and M.M. Jowkar. 2013. AMMI analysis of wheat substitution lines for detecting genes controlling adaptability. Int. J. Adv. Biol. Biomed. Res. 1(9): 1112-1123.
Annicchiarico, P. and M. Perenzin. 1994. Adaptation Patterns and Definition of Macro-environments for Selection and Recommendation of Common-wheat Genotypes in Italy. Plant Breed. 113:197-205.
Badr, A., K. Muller, R. Schafer-Pregl, H. El Rabey, S. Effgen, H.H. Ibrahim, C. Pozzi, W. Rohde and F. Salamini. 2000. On the origin and domestication history of barley (Hordeum vulgare). Mol. Biol. Evol.17(4):499–510.
Bedada, G., A. Westerbergh, E. Nevo, A. Korol and K. J. Schmid. 2014. DNA sequence variation of wild barley Hordeum spontaneum (L.) across environmental gradients in Israel. Heredity. 112: 646-655.
Belachew, K. Y., N. H. Maina, W. M. Dersseh, B. Zeleke and F. L. Stoddard. 2022. Yield gaps of major cereal and grain legume crops in Ethiopia: A review. Agronomy.12: 2528.
Chapagain, T. and A. Good. 2015. Yield and production gaps in rainfed wheat, barley, and canola in Alberta. Front. Plant Sci. 6:990.
Cossani, C. M., G. A. Slafer and R. Savin. 2010. Co-limitation of nitrogen and water, and yield and resource-use efficiencies of wheat and barley. Crop Past. Sci. 61: 844-851.
Díaz, M. J., M. Moya and E. Castro. 2022. Bioethanol production from steam-exploded barley straw by co-fermentation with Escherichia coli SL100. Agronomy. 12: 874.
Ebdon, J.S. and H.G. Gauch. 2002. Additive main effect and multiplicative interaction analysis of national turfgrass performance trials: I. Interpretation of genotype × environment interaction. Crop Sci. 42: 489-496.
Ebrahimnejad, S. and H. Sabouri. 2018. Evaluation of genotype× interaction effects on grain yield of barely genotypes using additive main effects and multiplicative interactions (AMMI). J. Crop Breed. 9: 144-151.
Erdemci, I., 2018. Investigation of genotype× environment interaction in chickpea genotypes using AMMI and GGE biplot analysis. Turkish Journal of Field Crops 23: 20-26.
Fana, G., D. Tadese, H. Sebsibe and R. P. Verma, 2018: Multi-environment trial analysis of food barley in Ethiopia using AMMI and GGE biplot methods. J. Plant Breed. Genet. 6: 75-85.
FAOSTAT. 2020. Food and Agriculture Organization of the United Nations, Rome, Italy. (www.fao.org/faostat/en/#data)
Gauch, H. G. 1992. Statistical analysis of regional yield trials: AMMI analysis of factorial designs. Elsevier, Amsterdam, Netherlands.
Gauch, H. G. and R.W. Zobel. 1996. AMMI Analysis of Yield Trials. In: Genotype-by-environment Interaction, Eds. Kang, M. S. and Gauch, H. G. CRC Press, Boca Raton, FL. p. 85-122.
Homma, S. 2015. AMMI, Stability and GGE biplot analysis of durum wheat grain yield for genotypes tested under some optimum and high moisture areas of Ethiopia. Acad. J. Entomol. 8(3): 132-139.
Islam, M.R., M. Anisuzzaman, H. Khatun, N. Sharma, M.Z. Islam, A. Akter and P.S. Biswas. 2014. AMMI analysis of yield performance and stability of rice genotypes across different Haor areas. Eco-Friendly Agric. J. 7(02): 20-24.
Kaur, V., J. Aravind, Manju, S. R. Jacob, J. Kumari, B. S. Panwar, N. Pal, J. C. Rana, A. Pandey and A. Kumar. 2022. Phenotypic characterization, genetic diversity assessment in 6,778 accessions of barley (Hordeum vulgare L. ssp. vulgare) germplasm conserved in National Genebank of India and development of a core set. Front. Plant Sci. 13: 771920.
Kebede B, A. and A. Getahun. 2017. Adaptability and stability analysis of groundnut genotypes using AMMI model and GGE-biplot. J. Crop Sci. Biotechnol. 20: 343-349.
Lyu, Y., S. Ma, J. Liu and X. Wang. 2022: A systematic review of highland barley: Ingredients, health functions and applications. Grain Oil Sci. Technol. 5: 35-43.
Meints, B., C. Vallejos and P. Hayes. 2021. Multi-use naked barley: A new frontier. J. Cereal Sci. 102: 103370.
Mittal, S. 2022. Wheat and Barley Production Trends and Research Priorities: A Global Perspective. In: P. L. Kashyap, V. Gupta, O. Prakash Gupta, R. Sendhil, K. Gopalareddy, P. Jasrotia and G. P. Singh eds. New Horizons in Wheat and Barley Research: Global Trends, Breeding and Quality Enhancement. Springer Singapore, Singapore. p.3-18.
Mortazavianc, S.M.M, H.R. Nikkhah, F.A. Hassani, M. Sharif-al-Hosseini, M. Taheri and M. Mahlooji. 2014. GGE biplot and AMMI analysis of yield performance of barley genotypes across different environments in Iran. J. Agric. Sci. Technol.16: 609-622.
Newman, C. W., R. K. Newman and C. E. Fastnaught. 2019: Barley Whole Grains and their Bioactives.p. 135-167.
Obadi, M., J. Sun and B. Xu. 2021. Highland barley: Chemical composition, bioactive compounds, health effects, and applications. Food Res. Int. 140: 110065.
Perera, W. N. U., M. R. Abdollahi, F. Zaefarian, T. J. Wester and V. Ravindran. 2022. Barley, an undervalued cereal for poultry diets: Limitations and opportunities. Animals. 12: 2525.
Purchase, J. L., H. Hatting and C.S. Vandeventer. 2000. Genotype × environment interaction of winter wheat (Triticum aestivum L.) in South Africa: II. Stability analysis of yield performance. S. Afr. J. Plant Soil. 17:101–107.
Raud, M., L. Rocha-Meneses, D. J. Lane, O. Sippula, N. J. Shurpali and T. Kikas. 2021. Utilization of barley straw as feedstock for the production of different energy vectors. Processes. 9: 726.
Russell, J., M. Mascher, I. K. Dawson, S. Kyriakidis, C. Calixto, F. Freund, M. Bayer, I. Milne, T. Marshall-Griffiths, S. Heinen et al. 2016. Exome sequencing of geographically diverse barley landraces and wild relatives gives insights into environmental adaptation. Nat. Genet. 48: 1024-1030.
Sakellariou, M. and P. V. Mylona. 2020. New uses for traditional crops: The case of barley biofortification. Agronomy. 10: 1964.
Schils, R., J. E. Olesen, K.-C. Kersebaum, B. Rijk, M. Oberforster, V. Kalyada, M. Khitrykau, A. Gobin, H. Kirchev, V. Manolova, et al. 2018. Cereal yield gaps across Europe. Eur. J. Agron. 101: 109-120.
Sharma, R., S. Mokhtari, S. M. Jafari and S. Sharma, 2022. Barley-based probiotic food mixture: health effects and future prospects. Crit. Rev. Food Sci. Nutrit. 62: 7961-7975.
Shukla, S., B.K. Mirshra, A. Siddiqui, R. Pandey and A. Rastogi. 2015. Comparative study for stability and adaptability through different models in developed high thebaine lines of opium poppy (Papaver somniferum L.). Ind. Crop. Prod. 74:875–886.
Soufan, W. and N. A. Al-Suhaibani. 2021. Optimizing yield and quality of silage and hay for pea-barley mixtures ratio under irrigated arid environments. Sustainability. 13: 13621.
Tricase, C., V. Amicarelli, E. Lamonaca and R. L. Rana. 2018. Economic analysis of the barley market and related uses. Grasses as Food and Feed. p.10.
Tse, T. J., D. J. Wiens, J. Shen, A. D. Beattie and M. J. T. Reaney. 2021. Saccharomyces cerevisiae fermentation of 28 barley and 12 oat cultivars. Fermentation. 7: 59.
Ullrich, S. E. 2014. The Barley Crop: Origin and Taxonomy. Barley: Chemistry and Technology, 1.
UnNisa, Z., A. Govind, M. Marchetti and B. Lasserre. 2022. A review of crop water productivity in the Mediterranean basin under a changing climate: Wheat and barley as test cases. Irrig. Drain. 71: 51-70.
USDA. 2018. Afghanistan: Low Precipitation Results in a Decline in Northern Winter Grains. Commodity Intelligence Report Production. (https://ipad.fas.usda.govhighlights201805Afghanistanindex.pdf).
Vaezi, B., A. Pour-Aboughadareh, R. Mohammadi, M. Armion, A. Mehraban, T. Hossein-Pour and M. Dorii. 2017. GGE Biplot and AMMI analysis of barley yield performance in Iran. Cereal Res. Commun. 45(3): 500–511.
VSN International. 2015. The Guide to the Genstat Command Language (Release 18), Part 2 Statistics. VSN International, Hemel Hempstead, UK.
Yirgu, M., M. Kebede, T. Feyissa, B. Lakew and A. B. Woldeyohannes. 2022. Morphological variations of qualitative traits of barley (Hordeum vulgare L.) accessions in Ethiopia. Heliyon. 8: e10949.
Zhang, W., G. Cao, X. Li, H. Zhang, C. Wang, Q. Liu, X. Chen, Z. Cui, J. Shen, R. Jiang, G. Mi, Y. Miao, F. Zhang and Z. Dou. 2016: Closing yield gaps in China by empowering smallholder farmers. Nature. 537: 671-674.
Zhang, W., J. Hu, Y. Yang and Y. Lin. 2020. One compound approach combining factor-analytic model with AMMI and GGE biplot to improve multi-environment trials analysis. J. Forest. Res. 31: 123-130.
Zhou, M.X. 2010. Barley Production and Consumption. In: G. Zhang and C. Li eds. Genetics and Improvement of Barley Malt Quality. Springer Berlin Heidelberg, Berlin, Heidelberg. p.1-17.
Zobel, R. W., M.J. Wright and H.G. Gauch. 1988. Statistical analysis of a yield trial. Agron. J. 80: 388–39.
Zohary, D. 2017. The progenitors of wheat and barley in relation to domestication and agricultural dispersal in the Old World. Domestication and exploitation of plants and animals. Routledge. p. 47-66.

Copyright © Akbarzai et al., 2022. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium provided the original author and source are appropriately cited and credited.
Join Journal of Environmental and Agricultural Sciences (JEAS)
Interested to join the JEAS Team
Join JEAS as a member Editorial Board see Editors’ Responsibilities
Join JEAS as a member Review Panel Reviewers’ Responsibilities
(send your CV through email at editor.jeas@outlook.com)
JEAS Indexing Journal of Environmental EAS is indexed by reputed indexing services.
Suggest Indexing service/s through email (editor.jeas@outlook.com)
Call for Articles
Submit Your research for publication in the “Journal of Environmental and Agricultural Sciences (JEAS)” through email: editor.jeas@outlook.com
- How to prepare your manuscript before submission
- How to submit your manuscript
- Publication Ethics
- Publication Fee Currently JEAS is publishing manuscripts without publication or processing fee
JEAS Recently Published and Highly Cited Articles
Citation record of JEAS: JEAS Google Scholar page
Follow JEAS Facebook
